EoE Pathophysiology
Great strides have been made in delineating the molecular events that lead to eosinophilic esophagitis (EoE), but a complete picture has yet to be created. Development of EoE is based on the complex interrelationship between the food antigens/luminal environment, the esophageal mucosa, and the immune system.1 In patients with EoE, as many as 80% have a history of atopy, including food allergy.2 Similar to atopic diseases, such as asthma and atopic dermatitis, the immune system involvement in EoE is polarized for type 2 inflammatory response.1 There is also a genetic component, with close family members more likely to develop the disease.3
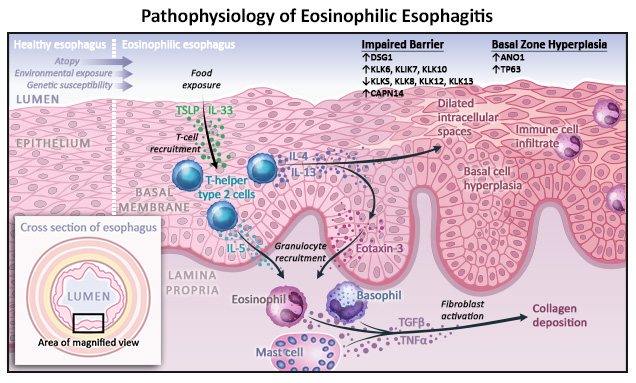
It is helpful to magnify and examine elements of EoE pathophysiology in a stepwise fashion, beginning with ingestion and movement of food antigens and/or microorganisms through the esophagus. However, the existence of an initiating trigger for EoE pathological processes, and which portion of the disease mechanism that might be, remains unclear and is currently under investigation. Elements of EoE development include (Figure 1):
- Epithelial interaction. Normally, the esophageal epithelium creates a strong barrier to keep luminal contents from gaining access to the bloodstream and tissues.4 In these layers, the apical junction complex with tight junctions, adherens junctions, and desmosomes maintains the barrier function and regulates the intracellular space between epithelial cells.
- In EoE, however, epithelial cells and dendritic cells in the mucosa come into contact with luminal antigens and release alarmins, which signal tissue damage, T-helper type 2 (Th2) cells to the area, and stimulate tissue-resident type 2 innate lymphoid cells (ILC2s) to expand and remain in tissues.2,5 Additionally, dendritic cells present antigens on their cell surface to further stimulate adaptive immunity.1,3 Th2 and ILC2s in the esophageal epithelia release the key Th2 cytokines, interleukin (IL)-4, IL-13, and IL-5.5
- Eosinophil maturation, activation, and survival are enhanced in response to IL-5 release. The release of IL-4/IL-13 into the environment leads to the stabilization of mast cells and an increase in vascular endothelial adhesion strength.1,3 Further, IL-4/IL-13 release enhances expression of the genes responsible for proliferation, inflammation, and barrier function.3
- The action of cytokines on immune cells occurs via signaling inside the cell by Janus kinase/signal transducer and activator of transcription (JAK/STAT) signaling proteins. After cytokines bind their receptors, JAK proteins bound to the receptor tail are activated, in turn activating their STAT binding partner to move into the nucleus alter genetic expression.6
- Changes in genetic expression that occur in response to Th2/ILC2 cytokine release lead to the remodeling of the esophageal mucosa. Cells in the basal zone of the epithelium become hyperplastic and epithelial intracellular spaces dilate.3 Dilation occurs due to impaired function of the tight junctions, adherens junctions, and desmosomes.4 Further, mast cells in the mucosa express transforming growth factor (TGF)-β, stimulating fibroblasts to lay down extracellular matrix, including collagen and fibronectin deposition.3 Over time, the increase in extracellular matrix components leads to the strictures associated with later stages of EoE.
References
- Muir A, Falk GW. Eosinophilic esophagitis: A review. JAMA. 2021;326(13):1310-1318. doi:10.1001/jama.2021.14920
- Racca F, Pellegatta G, Cataldo G, et al. Type 2 inflammation in eosinophilic esophagitis: From pathophysiology to therapeutic targets. Frontiers Physiol. 2022:12(815842):1-30. doi:10.3389/fphys.2021.815842
- Khokhar D, Marella S, Idelman G, et al. Eosinophilic esophagitis: Immune mechanisms and therapeutic targets [published online ahead of print, 2022 Jul 1]. Clin Exp Allergy. 2022;10.1111/cea.14196. doi:10.1111/cea.14196
- Blevins CH, Iyer PG, Vela MF, Katzka DA. The esophageal epithelial barrier in health and disease. Clin Gasteroenterol Hepatol. 2018;16:608-617. doi:10.1016/j.cgh.2017.06.035
- Wang YM, Bakhtiar M, Alexander SO. ILC2: There’s a new cell in town. J Am Soc Nephrol. 2017;28(7):1953-1955. doi:10.1681/ASN.2017040398
- Hu X, Ii J Fu M, Zhao X, Wang W. The JAK/STAT signaling pathway: From bench to clinic. Signal Transduct Target Ther. 2021;6(1):402. doi:10.1038/s41392-021-00791-1